Back in October 2019, we took a look at performance and file sizes for a handful of binary file formats for storing data frames in Python and R. These included Apache Parquet, Feather, and FST.
In the intervening months, we have developed “Feather V2”, an evolved version of the Feather format with compression support and complete coverage for Arrow data types. In this post, we explain what Feather V2 is and what you might find it useful. We also revisit the benchmarks from six months ago to show how compressed Feather V2 files compare, demonstrating that they can be even faster than Parquet to read and write. We also discuss some of the situations in which using Parquet or Feather may make more sense.
More about Feather V2
Wes and Hadley developed the original Feather format (“Feather V1”) early in 2016 as a proof of concept of using Arrow Arrow for fast, interoperable frame storage. While it has been extremely useful for people these last 4 years as a fast and simple way to move data between R and Python, it had a few shortcomings:
- No support for list columns
- No support for incremental / chunk-based writes
- Memory limitations: individual string columns limited to 2 gigabytes.
The “Arrow IPC format”, part of the Arrow columnar specification, is designed for transporting large quantities of data in chunks and does not have these limitations. IPC stands for “interprocess communication”, and we use this term to describe the format as it is designed for moving datasets from one process (or machine) to another.
We layered simple compression using the industry standard LZ4 or ZSTD libraries on top of the Arrow IPC format to create Feather V2. The result is files that are substantially smaller (but generally not as small as Parquet files, more on this below) but very fast to read and write. We chose to use LZ4 and ZSTD because they have extremely fast decompression speeds, substantially faster than zlib, Snappy, and other compression libraries in wide use.
Additionally, Feather V2 supports incremental and chunked writes. This allows you to build a large file without having to first load all of the data into memory. We have a little work to do to expose this functionality publicly (see ARROW-8470) but the format supports it.
Lastly, by basing Feather V2 on the Arrow IPC format, we assure longer-term stability of the file storage, since Apache Arrow has committed itself to not making backwards incompatible changes to the IPC format / protocol.
Benchmarking
As in the last performance analysis, we’re looking at two large (> 1 Gigabyte) datasets containing a mix of numeric and string data:
- Fannie Mae Loan Performance. We use the 2016Q4 “Performance” dataset, which is a 1.52 GB uncompressed CSV and 208 MB when gzipped
- NYC Yellow Taxi Trip Data: We use the “January 2010 Yellow Taxi Trip Records,” which is a 2.54 GB uncompressed CSV
For each dataset, we look at the following storage variants:
- Parquet, both with Snappy-compressed and Uncompressed internal data pages. Note that Parquet does a bunch of other encoding beyond using compression libraries
- Feather V2 with Uncompressed, LZ4, and ZSTD (level 1), and Feather V1 from
the current
featherpackage on CRAN - R’s native serialization format, RDS
- FST format with
compress = 0andcompress = 50(default)
For each case we compute:
- Read and write time to/from
pandas.DataFrame(in Python) anddata.frame(in R) - Read and write time from Arrow columnar table format (identical in Python and R)
- The size of each file
Methodology notes:
- CPU: i9-9880H laptop with 8 physical cores and a fast M.2 NVMe SSD. Many thanks to Dell for donating this hardware to Ursa Labs!
- We test performance using 1, 4, and 8 cores. Because the observed difference between using 4 and 8 cores was small relative to the difference between 1 and 4, the 8-core results are not shown in the plots in order to make them easier to read.
- Ubuntu 18.04 with CPU governor set to “performance”
- We use 0.17.0 release of Apache Arrow, which was just released. Python 3.7 and R 3.6 are used. The benchmarking scripts (a bit messy, sorry) are located on GitHub. If we made any mistakes, please let us know and we will fix it!
- For the read benchmarks, we read the file 5 times and compute the average of the 5 runs. By reading the file many times we hopefully partially mitigate the impact of disk caching in the Linux kernel, but this could be improved. For the write benchmarks we timed based on 3 iterations.
- For the Feather benchmarks that read Arrow tables, we disabled memory mapping so that we are aren’t “cheating” by using the zero-copy Arrow read functionality. That said, if you are working directly with Arrow tables, then uncompressed Feather files are effectively “free” to read (aside from the cost of disk IO) because of Arrow’s zero-copy memory-mapping functionality.
Results
For simplicity, the graphs below show benchmarks on the Fannie Mae dataset using 4 cores. The patterns observed are consistent with the NYC Taxi dataset, and four cores are generally better than one. If you’re interested in comparing, full benchmarks for read and write performance across both datasets are also posted, though they’re harder to read.
File size
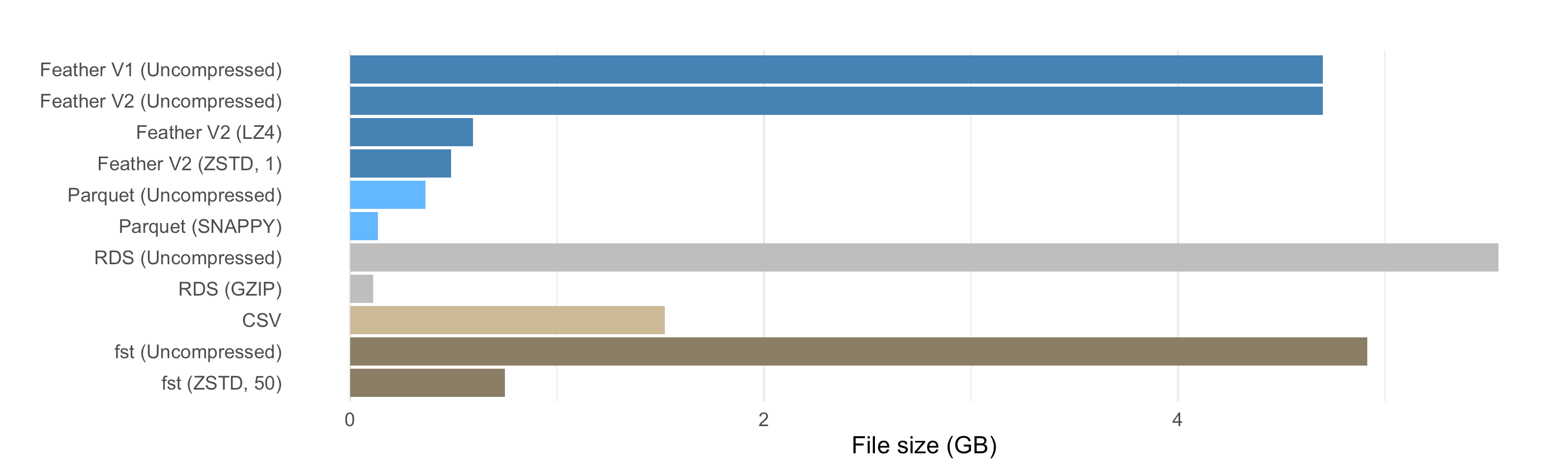
The file size benefits of compression in Feather V2 are quite good, though Parquet is smaller on disk, due in part to its internal use of dictionary and run-length encoding. Even without adding Snappy compression, the Parquet file is smaller than the compressed Feather V2 and FST files. Note that LZ4 and ZSTD have been added to the Parquet format but we didn’t use them in the benchmarks because support for them is not widely deployed.
Compressed .rds files (using GZIP) from R are slightly smaller than the Snappy-compressed Parquet files, but that comes at a cost, as we’ll see in the read and write speed results below. (And of course, they’re only readable by R.)
The relative file sizes for the nyc-taxi dataset are consistent with the patterns shown here, though due to the differences in data types, the benefit of compression is smaller across the board.
Read speed
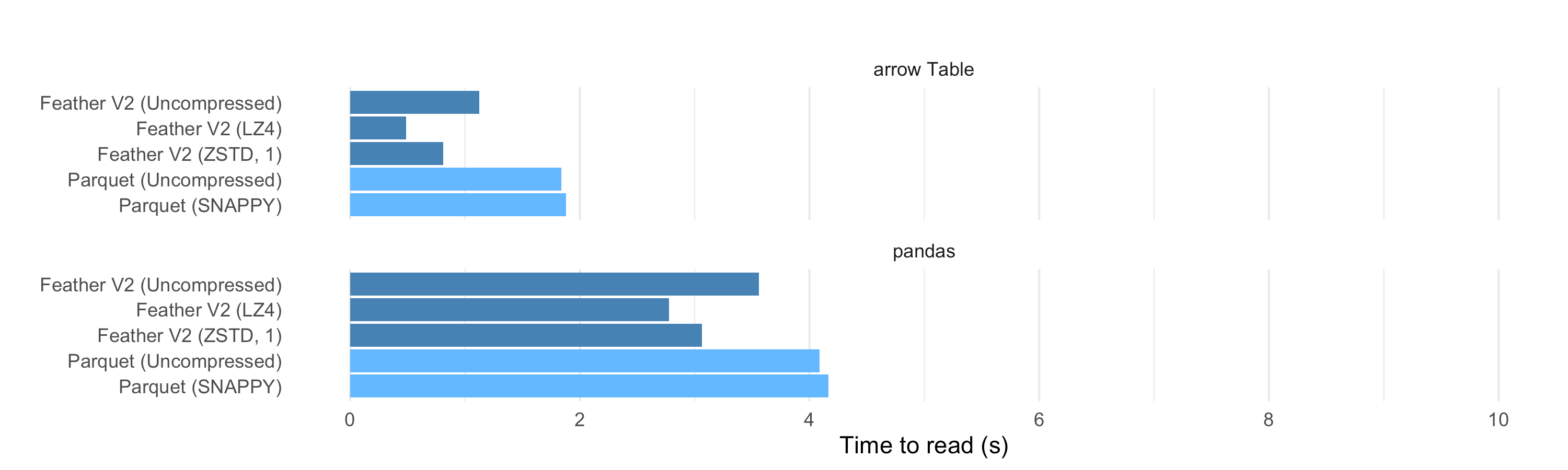
Overall, the read performance of the binary formats falls in the same general ballpark, and we see modest performance improvements by using multiple threads, which helps both with file decoding as well as the conversions to and from Arrow format.
Parquet is fast to read into an Arrow Table, but Feather is faster. And Feather with compression is even faster to read. This is partly because the files are much smaller on disk, so even though the NVMe SSD reads at 1-2GB/second, it shows that IO is a bottleneck.
For Parquet and Feather, performance of reading to Pandas and R is the speed of reading to Arrow plus the speed of converting that Table to a Pandas/R Data Frame. For the Pandas with the Fannie Mae dataset, we see that Arrow to Pandas adds around 2 seconds to each read. In our last performance analysis, we demonstrated that there is significant overhead associated with converting to and from Python pandas and R data frame objects. While indeed we’ve done our best to optimize the creation of these objects, populating data frames with a lot of strings is a bottleneck in both languages. As such, the read and write times of these file formats can be overshadowed by the effort involved with creating the data frames.
Some might comment that faster readers for these file formats could be built that circumvent the Arrow columnar format as the “middle man”. In the case of Feather, the stored data is Arrow format already, so there’s no extra overhead there. With Parquet, we are decoding Parquet files into Arrow first then converting to R or pandas. so strictly speaking faster “custom” converters could potentially be created, but we would guess the performance gains would be measly (at most 20%) and so hardly justify the implementation and code maintenance effort.
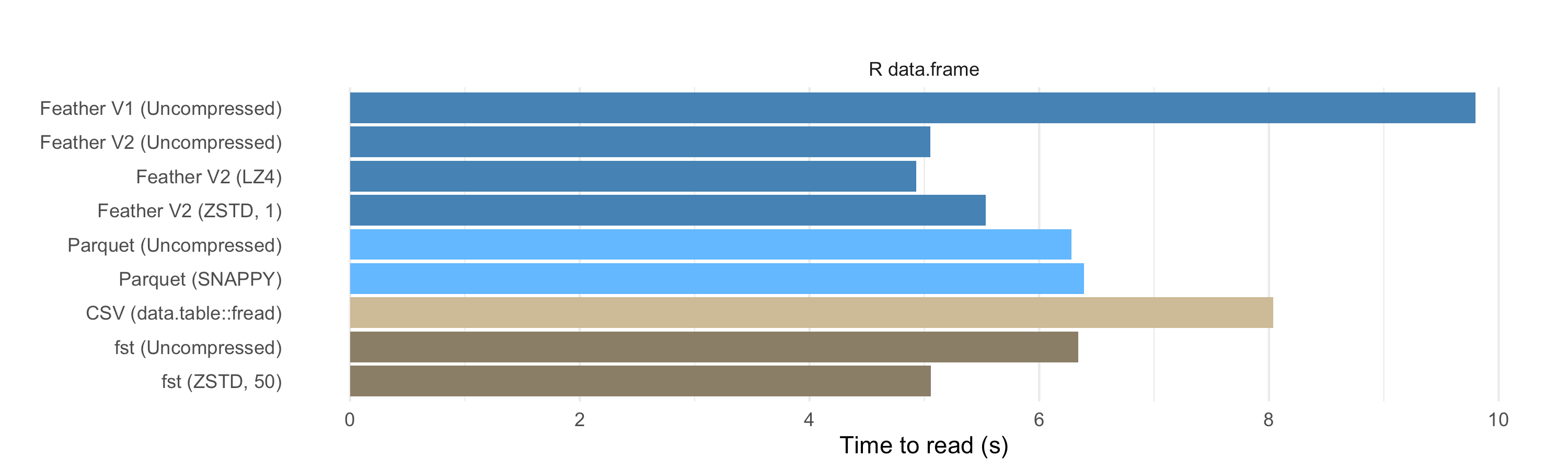
When reading in R, we see that the Arrow to R code is not as fast as the Arrow
to Pandas code. The Feather and Parquet R readers are about 2 seconds slower
than their Python counterparts. In the full results,
we can see that the timings for single-threaded Feather and Parquet reading in
Python and R are closer together. This may mean that the Arrow-to-R code isn’t
able to take as full advantage of multithreading compared with the similar code
in pyarrow, so there may be room for improvement there.
As we showed previously, the arrow package’s Feather reader (V2) is much
faster than the V1 implementation in the feather package. Parquet, Feather
V2, and FST are all comparably fast. Reading .rds is much slower, literally off
the chart here: the uncompressed file took 30 seconds to read, and 41 seconds
to read the compressed file. This highlights, among other things, that while
GZIP can achieve good compression ratios (i.e. smaller files), it is not as
fast as LZ4 or ZSTD, which is why Feather supports those two compression
libraries. It also shows the benefit of multithreaded reading as the .rds
reader is single-threaded.
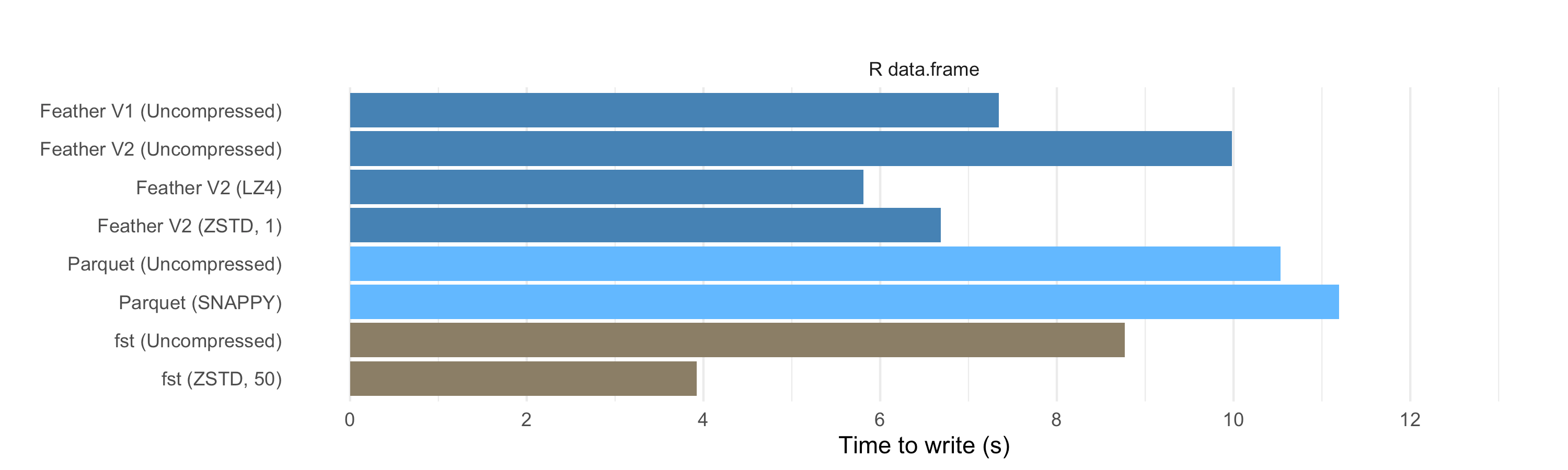
Write speed
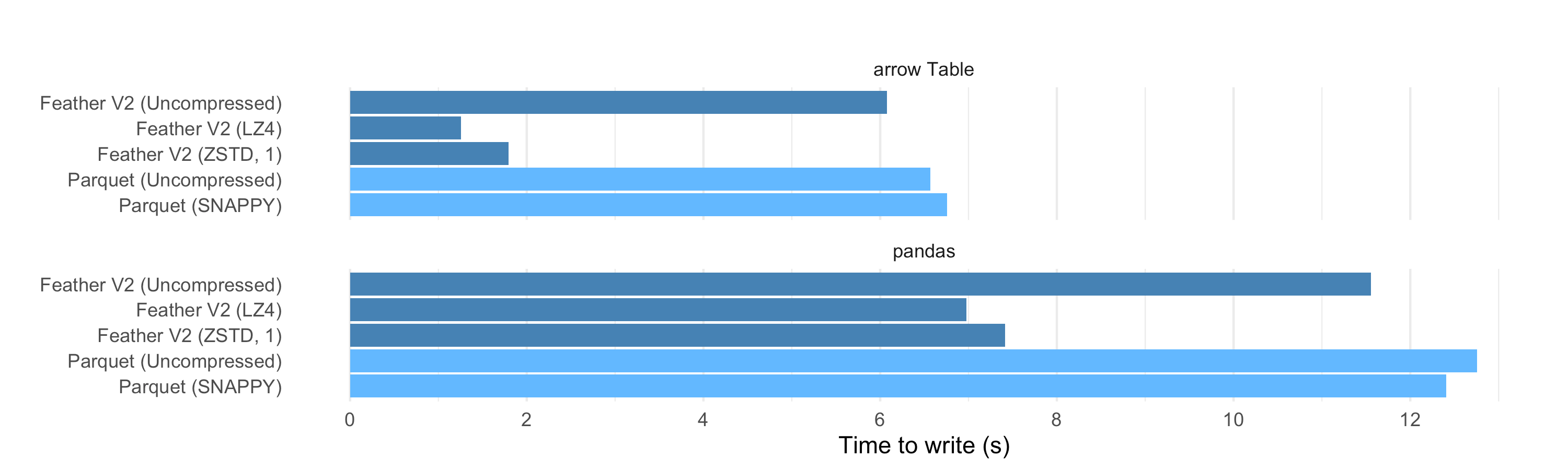
On the write side, we see that compressed Feather files can also be faster to write than uncompressed files, and also faster than writing to Parquet. So we can produce the compressed data at a higher speed than it would take to write the uncompressed data to disk, even with a very fast SSD. This is a significant performance win.
Why use Feather V2 when you can use Parquet?
Parquet format has become one of the “gold standard” binary file formats for data warehousing. We highly recommend it as a portable format that can be used in many different data processing systems. It also generally (but not always) produces very small files.
Feather V2 has some attributes that can make it attractive:
- Accessible by any Arrow implementation. R and Python use the Arrow C++ library internally, which is a well-supported reference implementation.
- Generally faster read and write performance when used with solid state drives, due to simpler compression scheme. When read over the network, we expect Parquet will outperform.
- Internal structure supports random access and slicing from the middle. This also means that you can read a large file chunk by chunk without having to pull the whole thing into memory.
- Complete support for all Arrow data types. Parquet has a smaller type system and while most of Arrow can be stored in Parquet, there are some things that cannot (like unions). Support for reading nested Parquet data is not fully implemented yet, but should be in 2020 (see ARROW-1644).
In general, our view is that having options are good, and you can choose the tool that works best for you.
Conclusion and Acknowledgements
We refer you to the October 2019 post on this topic for more information about some of the internal details relating to performance, especially with string-heavy datasets.
We hope again that this analysis is useful for Python and R programmers, and look forward to continuing to work with the open source community to make data processing faster and more productive!
This work was supported by some of our generous sponsors:
- RStudio
- Intel
- NVIDIA
- Two Sigma
- Bloomberg
If you would like to help fund Arrow development, please drop us a line at info@ursalabs.org.