Since the founding of Ursa Labs in 2018, one of our focus areas has been accelerating data access to binary file formats for Python and R programmers. My first semi-formal collaboration with the R community was the creation of the Feather file format with Hadley Wickham shortly after the Apache Arrow project was started. Since then I have focused my energy on the Arrow project to build high performance data processing infrastructure.
We have a number of motivations in working through Apache Arrow to accelerate data access:
- Take advantage of the memory- and CPU-efficient Arrow columnar memory layout to speed up processing especially of non-numeric (e.g. string) data
- Utilize a common C++ codebase in both Python and R and maintain that codebase with a large community of developers. We’ve had active collaborations with the Ruby and MATLAB on this codebase, which has been a lot of fun.
In August 2019, the R community published Apache Arrow to CRAN. Since the project is about to make its 0.15.0 release with a number of performance optimizations relating to Parquet files, we wanted to do some simple benchmarking to show how reading popular file formats performs in various scenarios.
File formats
The four columnar formats I look at are:
- Apache Parquet (simply “Parquet” from now on), a popular open standard columnar file format used widely in data warehousing. It began originally in the Apache Hadoop ecosystem but has been widely adopted by Apache Spark and by the cloud vendors (Amazon, Google, and Microsoft).
- Feather: a simple columnar format that Hadley and I created that can be described as “putting Arrow on disk”. Compared with Parquet it does not feature any encoding or compression; the Arrow data structures are written to disk unmodified, so a Feather file occupies the same amount of space on disk as it does in-memory
- FST: a binary columnar format designed for use in R
- RDS: R’s native serialization format
Note that out of the four of these, only Parquet and RDS support nested data (i.e. arrays, structs, etc.). The benchmarks in this post, therefore, only deal with flat / non-nested data.
Since we’re evaluating read performance, we’ll also compare reading the same
data from CSV with data.table::fread(), even though CSV is not a columnar
file format.
Example Datasets and Methodology
I look at two large (> 1 Gigabyte) datasets containing a mix of numeric and string data:
- Fannie Mae Loan Performance. I use the 2016Q4 “Performance” dataset which is a 1.52 GB uncompressed CSV and 208 MB when gzipped
- NYC Yellow Taxi Trip Data: I use the “January 2010 Yellow Taxi Trip Records” which is a 2.54 GB uncompressed CSV
For each dataset, I generate files for each file format without any data preparation or cleansing. Some of these files would benefit from columns being converted to “dictionary-encoded” in Arrow directly, or equivalently “factor” in R or “categorical” in pandas, but I leave them as normal strings since this is more representative of what the average user will experience.
As far as hardware, I use the following machine:
- i9-9880H laptop with 8 physical cores. I will test performance using 1, 4, and 8 cores
- Ubuntu 18.04 with CPU governor set to “performance”
Many thanks to Dell for donating this hardware to Ursa Labs!
Version information:
- Python interpreter 3.7.4
- Python pyarrow 0.15.0
- Python pandas 0.24.2
- R interpreter 3.6.1
- R arrow 0.15.0
- R data.table 1.12.2
- R fst 0.9.0
Difference in file sizes
Before digging into the benchmark results and analysis, I want to point out the significant differences in on-disk file sizes of each of the file formats under consideration.
In the case of the Fannie Mae dataset, we have:
- 208 MB as gzipped CSV
- 503 MB as FST file
- 114 MB as Parquet (with dictionary-encoding and Snappy data page compression)
- 3.96 GB as a Feather file (due to the absence of any encoding or compression). Note that adding compression to Feather files would be a straightforward affair and we would be happy to accept a pull request for this
- 4.68 GB as an uncompressed R RDS file
That is, the Parquet file is half as big as even the gzipped CSV. One of the reasons that the Parquet file is so small is because of dictionary-encoding (also called “dictionary compression”). Dictionary compression can yield substantially better compression than using a general purpose bytes compressor like LZ4 or ZSTD (which are used in the FST format). Parquet was designed to produce very small files that are fast to read.
Benchmark results
I read the files in several different ways:
- Read Parquet to Arrow using
pyarrow.parquet - Read Parquet to Arrow using
pyarrow.parquetand convert topandas.DataFrame - Read Feather to pandas using
pyarrow.feather - Read Parquet to R
data.frameusingarrow::read_parquet - Read Feather to R
data.frameusingfeather::read_feather, the old implementation before we reimplemented Feather in Apache Arrow - Read Feather to R
data.frameusingarrow::read_feather, to show the performance improvements of thearrowpackage over thefeatherpackage - Read FST to R
data.frameusingfst::read_fst - Read CSV file with
data.table::fread - Read uncompressed R RDS file created with
saveRDS
The benchmarking scripts (a bit messy, sorry) are located on GitHub. If I made any mistakes, please let me know and I will fix it!
In each benchmark, I read the file 5 times and compute the average of the 5 runs. By reading the file many times I hopefully partially mitigate the impact of disk caching in the Linux kernel, but this could be improved.
Here are the performance results. The bars are color-coded by the type of data
structure created. Note that generating Apache Arrow columnar format is
significantly more efficient than either R data.frame or pandas.DataFrame. I
will discuss some reasons why this is below.
The results are split by dataset: see the facet titles on the right. The colors indicate what kind of object is produced at the end. On the left you can see each case as above. The measurement is the average read time; shorter bars indicate faster performance. When possible, the libraries use all available CPU cores to improve performance.
Note that these benchmarks were run with Apache Arrow 0.15.0, which includes some performance and memory use-related bug fixes to the previous 0.14.1 release.
Results with 8 threads (OMP_NUM_THREADS=8)
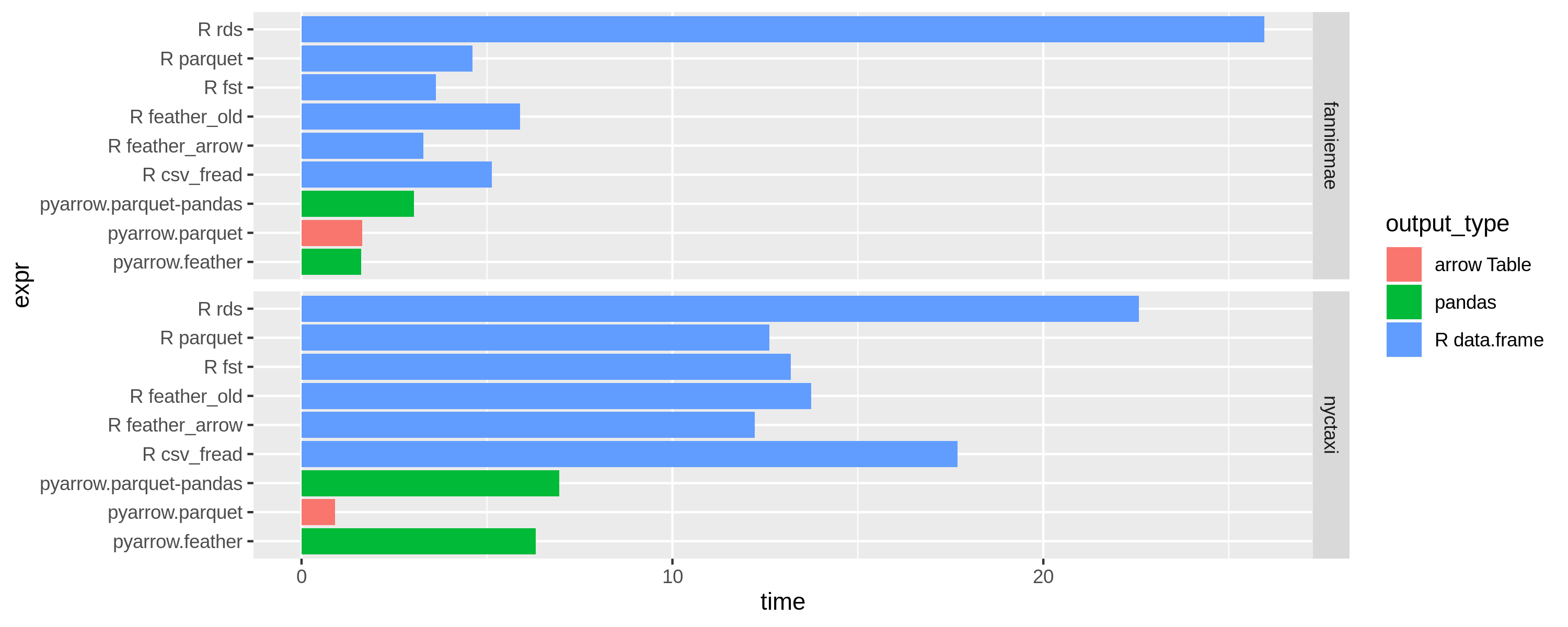
Results with 4 threads (OMP_NUM_THREADS=4)
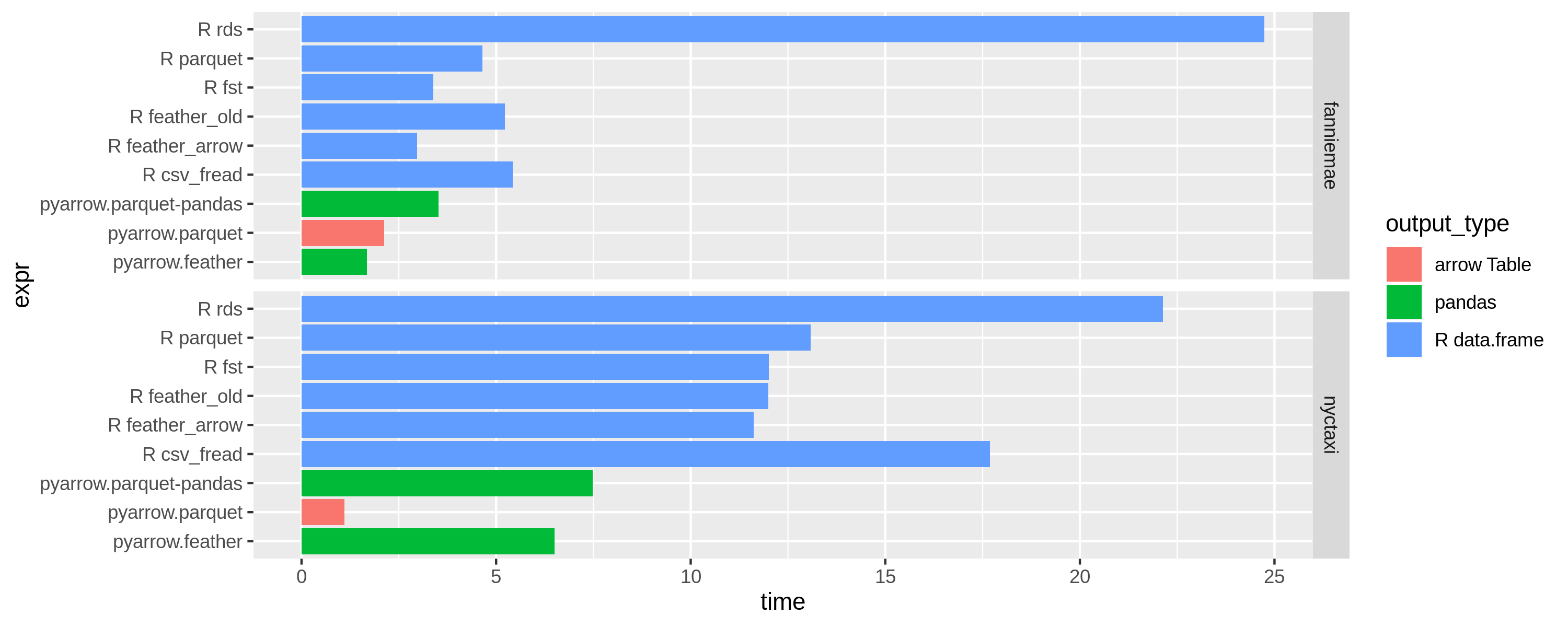
Results with 1 thread (OMP_NUM_THREADS=1)
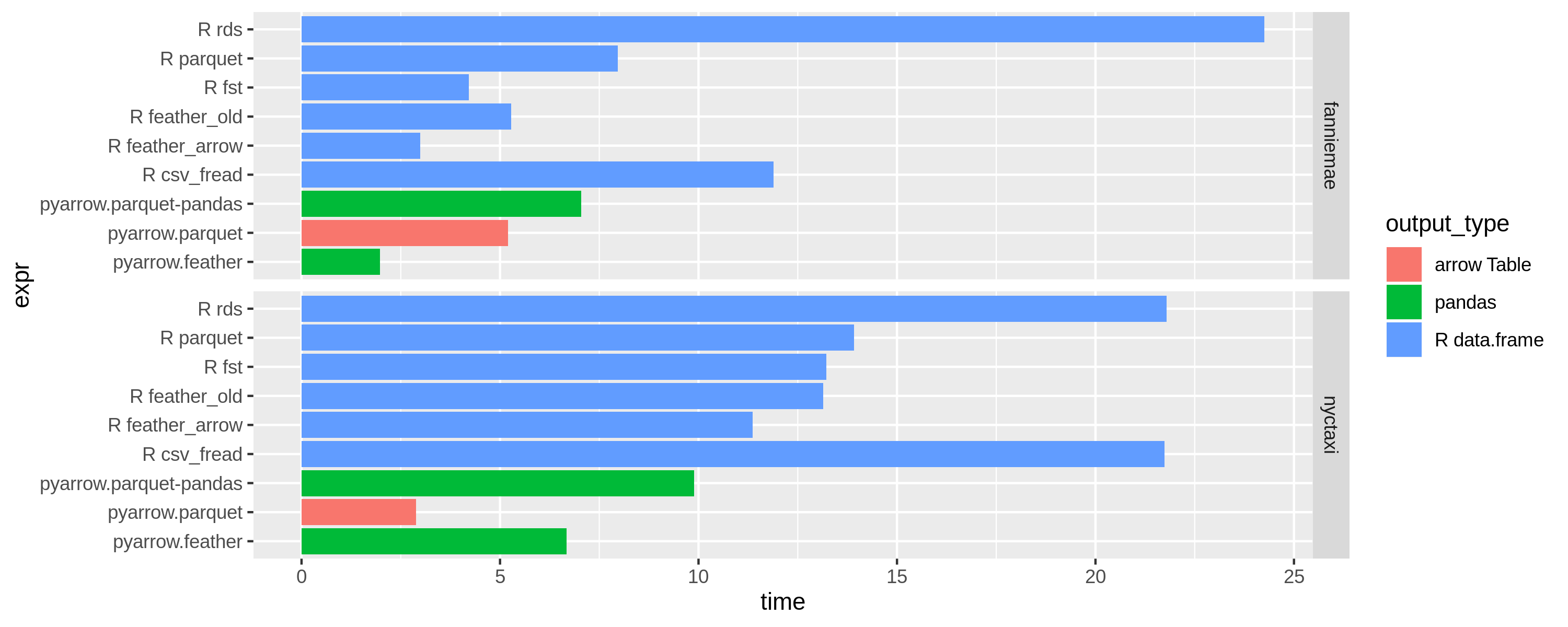
Performance result discussion
When controlling by output type (e.g. comparing all R data.frame outputs with
each other) we see the the performance of Parquet, Feather, and FST falls within a relatively small margin
of each other. The same is true of the pandas.DataFrame outputs.
data.table::fread is impressively competitive with the 1.5 GB file size but
lags the others on the 2.5 GB CSV.
There are a few things I would like to comment on:
- Why should you use Parquet over the other columnar formats?
- If you are a current user of the
featherR package, is it worth upgrading to thearrowpackage for dealing with Feather files? - Why is reading to Arrow format so much faster than to the other data frame formats?
- Why are the pandas benchmarks faster than the R benchmarks?
Parquet versus the other formats
Now that there is a well-supported Parquet implementation available for both Python and R, we recommend it as a “gold standard” columnar storage format. These benchmarks show that the performance of reading the Parquet format is similar to other “competing” formats, but comes with additional benefits:
- The size of Parquet files is significantly smaller in almost all cases, due to Parquet’s compression schemes
- Parquet is an industry-standard data format for data warehousing, so you can use Parquet files with Apache Spark and nearly any modern analytic SQL engine (Apache Impala, Google BigQuery, Presto, Amazon Redshift, etc.)
From feather to arrow in R
The arrow::read_feather function can be up to 50% faster than the
feather::read_feather function, particularly on multi-core CPU
environments. We recommend all feather users upgrade to use the arrow
package in R to take advantage of the optimization work we will continue to do
there.
Arrow versus the other output types
I used the Linux perf tool to investigate and found that
the main difference has to do with how strings are handled. As discussed in a
past Arrow blog post, string-heavy datasets are expensive to work with
when using pandas. This can be mitigated by using pandas.Categorical, but
this won’t necessarily help in data where the cardinality of the “unique set”
is high.
By contrast, the Apache Arrow string layout is extremely efficient. Each string value has a small amount of storage overhead (4 bytes plus 1 bit) and values are guaranteed to be adjacent to each other in memory, so the CPU won’t have as many “cache misses” when constructing or processing a string column.
I don’t know the exact details of how R deals with strings, but my understanding is that there is a “global string hash table” and so strings in R feature pointer indirection and likely a similarly high incidence of cache misses like pandas does. Note that because of R’s global hash table, string data in R in general may use less memory than in Python.
pandas.DataFrame vs. R data.frame
I haven’t done enough analysis to determine exactly where the performance difference between Python and R lies. Both implementations use the same Arrow C++ code to read the Parquet and Feather files into Arrow format. So the difference likely has to do with conversion of strings from Arrow into the other data frames' representations.
Compared with the Arrow R library, we have spent a significant amount of time
optimizing the performance of converting from C++ Arrow data structures to
pandas.DataFrame. I expect there are some performance gains to be found in
the Arrow-to-R conversion C++ code.
Conclusion and Acknowledgements
We hope this analysis is useful for Python and R programmers, and look forward to continuing to work with the open source community to make data processing faster and more productive!
This work was supported by some of our generous sponsors:
- RStudio
- NVIDIA
- Bloomberg
- Two Sigma
If you would like to help fund Arrow development, please drop us a line at info@ursalabs.org.